Human Exposure to Environmental Contaminants
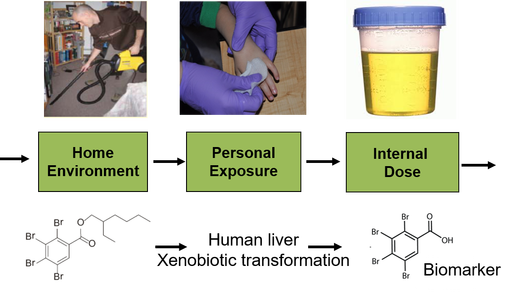
Environmental exposure plays a very important role in human health. In the real world, humans are exposed to “cocktails” of contaminants (e.g, exposome) and the characterization of all compounds is very challenging. In our lab, we have set up a sensitive analytical platform to analyze both routine and emerging contaminants in the environmental samples (indoor air, dust, water and products) and biological fluids (urine and blood) using mass spectrometry methods. These compounds include but are not limited to flame retardants, phthalates, dioxins, polychlorinated biphenyls, polycyclic aromatic hydrocarbons, pesticides, herbicides, plasticizers, and personal care products (PPCPs, e.g., triclosan and parabens) as well as their metabolites. Biomarker identification is a critical step in estimating human exposure. To investigate the biotransformation of some emerging compounds, both in vitro measurements with cell cultures and in vivo measurements will be conducted to examine metabolism. After identifying the primary metabolic pathways, biomonitoring of those chemicals and their metabolites in human biofluids (i.e. serum and urine) will be conducted and potential exposure biomarkers will be measured. Our current interest in human exposure field includes: (1) Xenobiotic transformation pathways of emerging organic pollutants; (2) Exposure biomarker identification for emerging organic pollutants; (3) Analytical platform development to analyze human exposome; (4) Correlation between environmental exposure and human health.
Metabolomics
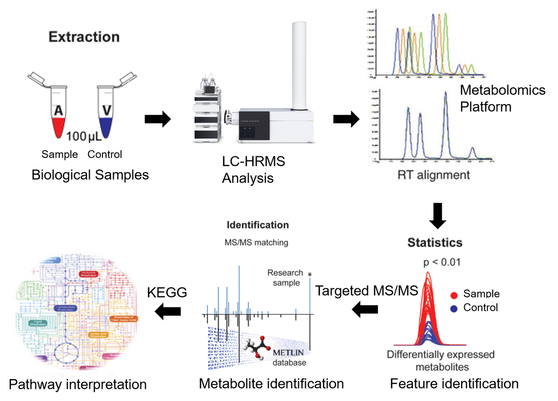
Metabolomics is the scientific study on the unique chemical fingerprints of specific cellular processes with a focus on small-molecule metabolite profiles. Together with proteomic and transcriptomic information, metabolomics provides a better understanding of cellular biology. Metabolomics has become increasingly important across a broad range of research areas including drug discovery, microbiology, plant physiology, and nutrition. Our group is interested in developing metabolomics platforms and their application. Our previous research focuses on the application of high-resolution mass spectrometry (HRMS)-based metabolomics to investigate metabolite and biological pathway perturbations in relation to disease conditions. Previous work include the identification of metabolism modulator on oligodendrocyte progenitor cell (OPCs) differentiation, and quantitative metabolomics on the effects of stem cell derived retinal pigment epithelium transplantation. Our recent publication has identified some key metabolites in modulating OPC differentiation (Beyer and Fang et al., 2017, Nature Chemical Biology, In Press). Our current interest on metabolomics can be summarized in following areas: (1) Metabolomics application in stem cell differentiation; (2) Metabolomics application in understanding the mode of action of environmental toxicants (primarily emerging organic contaminants); (3) Dynamic metabolic process during biological processes (Fluxomics); (4) Omics integration and its application on systems biology.
Xenobiotics and gut microbiota interaction
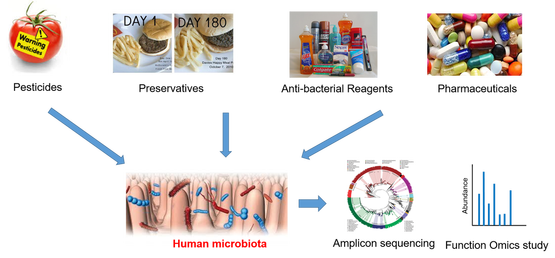
The gut microbiota is a complex ecosystem of microorganisms that colonize the gastrointestinal (GI) tract and its balance can play a variety of roles in the maintenance of human health condition. Accumulating evidence has also shown that the composition of microbiota could be affected and reshaped by environmental conditions. Human are ubiquitously exposed to synthetic chemicals produced during industrial and agricultural applications. The effects of those chemicals on human health are a global concern. Many of those EOCs including plasticizers, food preservatives and especially personal care products (PCPs) are designed to eliminate or hinder the growth of the microorganisms. Therefore, it is very possible that these compounds have biological effect on the gut microbiomes which in turn affect human health. Furthermore, the GI microbiome have broad enzymes which can metabolize environmental chemicals from various chemical families, modulating their toxicity to its host. Our research is to systematically investigate the biotransformation of EOCs by gut microbiomes and its effect on the metabolism of gut microbiomes at the environmentally relevant levels to address the following questions. Q1: How the contaminants would affect the metabolism of microbiota? Q2: How the microbiota would affect the fate of contaminants? Q3: What is the health effect of bacterial dysbiosis?
Bio-active compound screening from complex mixtures
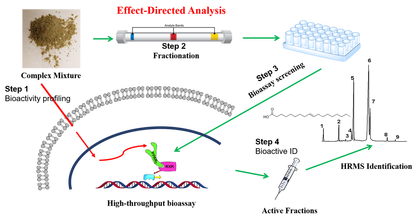
Bio-active compound screening from complex mixtures (e.g., crude extracts) is one of the most difficult and timely question for many pharmacologists and toxicologists. Effect-directed analysis (EDA) or bioassay directed analysis is a relatively recent technique utilizing iterative chemical separation, toxicity evaluation and qualitative chemical analysis to identify the chemical(s) exerting a specific toxicity in complex mixtures. In our previous studies, we have applied this technology to identify the causal compounds leading to the cardio-toxicity in zebra fish exposed to porewater from contaminated areas (Fang et al., 2014, Environmental Toxicology and Chemistry). Similarly, PPARgamma nuclear receptor agonists were found in human house dust samples (Fang et al., 2015, Environmental Science & Technology). Our current interest for this EDA can be summarized as follows: (1) Method development for fractionation techniques; (2) High-throughput bioassay development; (3) Non-known compound identification using high-resolution mass spectrometry (LC-HRMS and GC-HRMS); (4) Bio-active compound screening in the environmental (e.g, particulate matters or house dust) and gut microbial mixtures.
